-Search query
-Search result
Showing 1 - 50 of 8,823 items for (author: qi & y)

EMDB-44372: 
In-cell Saccharomyces cerevisiae nuclear pore complex with single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-44377: 
In-cell Saccharomyces cerevisiae nuclear pore complex with double nuclear ring and basket
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-44379: 
In-cell Mus musculus nuclear pore complex with nuclear basket
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-44381: 
In-cell Toxoplasma gondii nuclear pore complex
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E

EMDB-45197: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with double nuclear ring and basket
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45198: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45199: 
In-cell Saccharomyces cerevisiae nuclear pore complex cytoplasmic ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45200: 
In-cell Saccharomyces cerevisiae nuclear pore complex inner ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45201: 
In-cell Saccharomyces cerevisiae nuclear pore complex single nuclear ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45202: 
In-cell Saccharomyces cerevisiae nuclear pore complex double nuclear ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45203: 
In-cell Saccharomyces cerevisiae nuclear pore complex nuclear basket focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45204: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45205: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for double nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45216: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45219: 
In-cell Mus musculus nuclear pore complex cytoplasmic ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45220: 
In-cell Mus musculus nuclear pore complex inner ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45222: 
In-cell Mus musculus nuclear pore complex nuclear ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45223: 
In-cell Mus musculus nuclear pore complex basket focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45227: 
In-cell Mus musculus nuclear pore complex membrane focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45228: 
In-cell Toxoplasma gondii symmetry-expanded nuclear pore complex
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E
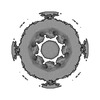
EMDB-45255: 
In-cell Saccharomyces cerevisiae C8-symmetrised nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45256: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45257: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45258: 
In-cell Mus musculus symmetry-expanded nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45259: 
In-cell Toxoplasma gondii C8-symmetrised nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E
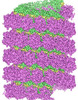

EMDB-31367: 
Structure of mumps virus nucleoprotein without C-arm
Method: helical / : Shen Q, Shan H, Zhang N, Qin Y

EMDB-39773: 
Cryo-EM structure of E.coli SPFH-NfeD family protein complex QmcA-YbbJ
Method: single particle / : Qiao Z, Gao YG

EMDB-38850: 
Cryo-EM structure of human dopamine transporter in apo state
Method: single particle / : Zhao Y, Li Y

EMDB-38851: 
Cryo-EM structure of human dopamine transporter in complex with dopamine
Method: single particle / : Zhao Y, Li Y

EMDB-38852: 
Cryo-EM structure of human dopamine transporter in complex with benztropine
Method: single particle / : Zhao Y, Li Y

EMDB-38853: 
structure of a proteinACryo-EM structure of human dopamine transporter in complex with GBR12909
Method: single particle / : Zhao Y, Li Y

EMDB-38854: 
Cryo-EM structure of human dopamine transporter in complex with methylphenidate
Method: single particle / : Zhao Y, Li Y

EMDB-38845: 
Icosahedrally averaged cryo-EM reconstruction of PhiKZ capsid before applying the "block-based" reconstruction method
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-38846: 
Block 1 of PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-38848: 
Block 2 of PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-39002: 
Composite cryo-EM map of PhiKZ capsid after applying the "block-based" reconstruction method
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

PDB-8y6v: 
Near-atomic structure of icosahedrally averaged jumbo bacteriophage PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-38695: 
Cryo-EM structure of ATP-DNA-MuB filaments
Method: helical / : Zhao X, Zhang K, Li S

EMDB-38696: 
CryoEM structure of ADP-DNA-MuB conformation1
Method: single particle / : Zhao X, Zhang K, Li S

EMDB-38697: 
CryoEM structure of ADP-DNA-MuB conformation2
Method: single particle / : Zhao X, Zhang K, Li S

EMDB-38698: 
CryoEM structure of ADP-DNA-MuB conformation3
Method: single particle / : Zhao X, Zhang K, Li S

EMDB-38699: 
CryoEM structure of ADP-DNA-MuB conformation4
Method: single particle / : Zhang X, Zhang K, Li S

PDB-8xvc: 
CryoEM structure of ADP-DNA-MuB conformation1
Method: single particle / : Zhao X, Zhang K, Li S

PDB-8xvd: 
CryoEM structure of ADP-DNA-MuB conformation2
Method: single particle / : Zhao X, Zhang K, Li S

EMDB-38300: 
Cryo-EM structure of partial dimeric WDR11-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39863: 
Cryo-EM structure of dimeric WDR11-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39943: 
Cryo-EM structure of dimeric WDR11-FAM91A1 complex Body2
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39947: 
Cryo-EM structure of dimeric WDR11-FAM91A1 complex Body1
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39949: 
Cryo-EM structure of WDR11-dm-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
